Abstract
Surface of an InAs wetting layer on GaAs(001) preceding InAs quantum dot (QD) formation was observed at 300°C with in situ scanning tunneling microscopy (STM). Domains of (1 × 3)/(2 × 3) and (2 × 4) surface reconstructions were located in the STM image. The density of each surface reconstruction domain was comparable to that of subsequently nucleated QD precursors. The distribution of the domains was statistically investigated in terms of spatial point patterns. It was found that the domains were distributed in an ordered pattern rather than a random pattern. It implied the possibility that QD nucleation sites are related to the surface reconstruction domains.
Similar content being viewed by others
Quantum dots (QDs) are potentially used for high-efficiency laser devices [1]. It is crucial to control QD formation to arrange QDs with high uniformity and high density. Little is known, however, of the growth mechanism of QDs, in particular the surface reconstruction of a wetting layer (WL) and QD nucleation sites in Stranski-Krastanow (S-K) mode. Because the surface reconstruction changes microscopically and dynamically in the course of WL growth, an in situ scanning tunneling microscopy (STM) technique such as STMBE [2] is essential. Atomic-scale in situ observation of an InAs WL on a GaAs(001) substrate has revealed that the surface reconstruction of the InAs WL changes from c(4 × 4) to the mixture of (1 × 3)/(2 × 3) and (2 × 4) prior to QD formation [3]. It is considered that such surface reconstructions form domains on InAs WL, and investigating their distribution will give a clue to understand a QD nucleation mechanism.
The distribution of reconstruction domains is characterized by spatial point patterns: a regular (ordered) pattern, a Poisson (random) pattern, and a clustered (aggregated) pattern [4]. In a regular pattern, points are distributed uniformly. Voronoi tessellation, that is a polygonal decomposition of a space by perpendicular bisector lines among neighboring points, is often used in spatial point analysis. The standard deviation of Voronoi cell areas represents well the point patterns. For more precise analysis, the distance to the nearest neighbor point from an arbitrary position, r 1, is helpful [5–7]. Let p(t) denote the probability that r 1 occurs less than t. The nearest neighbor distance function p(t) is identical to the probability of plotting a random point within the union area of circles whose radii are t and centers are the points. Trend of p(t) represents the characteristics of spatial point patterns.
In this paper, we investigate the surface reconstruction domains on InAs WL preceding QD formation by using in situ STM observation and discuss their distribution using spatial point analysis.
A piece (11 × 13 × 0.6 mm3) of GaAs(001) crystal was used as a substrate. First, the surface was thermally cleaned to remove the oxide layer under 1 × 10−4 Pa of an arsenic atmosphere in an MBE growth chamber. Next, a GaAs buffer layer was grown on the surface by using MBE until atomically smooth surface was obtained. The substrate was annealed at 430°C for 0.5 h to confirm the formation of c(4 × 4) reconstruction with reflection high-energy electron diffraction (RHEED). An STM unit was transferred to the sample holder in the growth chamber. A flux of In was irradiated to the sample during STM observation. After 1.5 monolayer (ML) of InAs WL growth, the substrate temperature was decreased to 300°C, and the As4 flux was shut off.
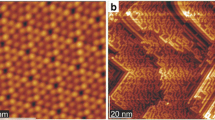
Figure 1 shows the STM image of 1.5 ML of InAs WL just prior to QD formation at 300°C. Stripes due to As dimers were clearly observed. The image was divided by a 25 × 25 mesh. The pitch of the As stripes, corresponding to the unit cell length along [110] azimuth of InAs surface reconstructions, was measured from the STM line profile for each cell. The data are plotted in the color diagram of Fig. 2. The pitch was classified into three ranges, namely the range from 0.6 to 1.0 nm, the range from 1.0 to 1.4 nm assuming (1 × 3)/(2 × 3), and the range from 1.4 to 2.0 nm assuming (2 × 4). Most of the cells had (1 × 3)/(2 × 3) or (2 × 4) surface reconstruction. Four neighboring cells having the same surface reconstruction were located in the diagram as indicated by oval markers in Fig. 3. A set of these cells correspond to a surface reconstruction domain extending for 16 nm2. For each of (1 × 3)/(2 × 3) and (2 × 4) surface reconstructions, the center points of the domains were marked, and their coordinates were measured by using ImageJ software [8, 9]. The center coordinates were used for the Voronoi tessellations of the STM view field (Fig. 4) and the computation of the nearest neighbor distance function p(t) [5]. The cells touching the frame of the STM image was not used for the computation since they are not true Voronoi cells. For the calculation of p(t), t was normalized by the factor f as follows:

where S is the total area of Voronoi cells, which are not touching the frame, and N is the number of valid reconstruction domains.
Pitch of arsenic dimer row for each cell of 25 × 25 mesh in Fig. 1
Voronoi tessellations of STM view field of Fig. 1 according to a (1 × 3)/(2 × 3) domains and b (2 × 4) domains
The density of each surface reconstruction domain is listed in Table 1. Both surface reconstruction domains had similar densities in the STM image of Fig. 1. Since these values were comparable to the typical density (∼1 × 1012 cm−2) of InAs QD precursors nucleating afterward, it implies the possibility that a QD formation pattern is based on the distribution of surface reconstruction domains [3].
The standard deviation of Voronoi cells for each surface reconstruction domain is also listed in Table 1. The total area of the Voronoi cells that are not touching the edge of the view field was normalized to 1.0 for the calculation. A typical value of a Poisson pattern by scattering 50 random points was ∼0.4, whereas that of the surface reconstruction domains was ∼0.3.
The nearest neighbor distance function p(t) of the surface reconstruction domains will give more precise information. Figure 5 shows the traces of p(t), which were calculated for the surface reconstruction domains as well as a typical regular point pattern. The p(t) envelope region of typical Poisson patterns was calculated by accumulating 50 simulations of scattering 50 random points. Traces of the surface reconstruction domains were plotted between that of the ordered pattern and the Poisson envelope region. This shows that the surface reconstruction domains were distributed in an ordered pattern rather than a random pattern. If we compare p(t) traces between surface reconstruction domains and QD precursors just after nucleation, the relationship between them and QD growth mechanism will be known more precisely.
In conclusion, (1 × 3)/(2 × 3) and (2 × 4) domains were located in the in situ STM image of 1.5 ML of InAs WL preceding QD nucleation. The densities of the reconstruction domains were similar to that of QD precursors just after nucleation. Spatial point analysis of the surface reconstruction domains revealed that the domains were distributed in an ordered pattern rather than a typical random pattern.
References
Arakawa Y, Sakaki H: Appl. Phys. Lett. 1982, 40: 939. COI number [1:CAS:528:DyaL38XkvVansrg%3D]; Bibcode number [1982ApPhL..40..939A] COI number [1:CAS:528:DyaL38XkvVansrg%3D]; Bibcode number [1982ApPhL..40..939A] 10.1063/1.92959
Tsukamoto S, Koguchi N: J. Cryst. Growth. 1999,201(202):118. Bibcode number [1999JCrGr.201..118T] Bibcode number [1999JCrGr.201..118T] 10.1016/S0022-0248(98)01304-9
Tsukamoto S, Honma T, Bell GR, Ishii A, Arakawa Y: Small. 2006, 2: 386. COI number [1:CAS:528:DC%2BD28Xhs1aqsLs%3D] COI number [1:CAS:528:DC%2BD28Xhs1aqsLs%3D] 10.1002/smll.200500339
Diggle PJ: Statistical Analysis of Spatial Point Patterns. Oxford University Press Inc., New York; 2003.
Tanemura M, Ogata Y: Suuri Kagaku (Math. Sci.). 1981, 213: 11. in Japanese in Japanese
Ogata Y, Tanemura M: Ann. Inst. Stat. Math.. 1981, 33: 315. 10.1007/BF02480944
Baddeley A, Gill RG: Ann. Stat.. 1997,25(1):263. 10.1214/aos/1034276629
Rasband WJ, :ImageJ. US National Institutes of Health, Bethesda, MD; 1997–2009. [http://rsb.info.nih.gov/ij/]
Abramoff MD, Magelhaes PJ, Ram SJ: Biophotonics Int. 2004,11(7):36.
Acknowledgments
Authors are grateful to Mr. Minoru Yamamoto, Ms. Sayo Yamamoto, and Mr. Hisanori Iwata.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Konishi, T., Tsukamoto, S. Statistical Analysis of Surface Reconstruction Domains on InAs Wetting Layer Preceding Quantum Dot Formation. Nanoscale Res Lett 5, 1901 (2010). https://doi.org/10.1007/s11671-010-9754-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11671-010-9754-3